Scientists have recently conducted a study, which has been published in the reputable journal PeerJ Life & Environment and featured in the International Association for Biological Oceanography Hub, to investigate the effectiveness of DNA metabarcoding in identifying fish eggs.
The primary objective of this research was to evaluate the performance of DNA metabarcoding, a process that enables the amplification and sequencing of a conserved gene found in all animals, in increasing the efficiency and reducing the costs of a long-term fish egg monitoring program.
According to the study, the results were quite promising. The fish egg identifications were consistent with prior species distributions observed from individual egg DNA barcoding.
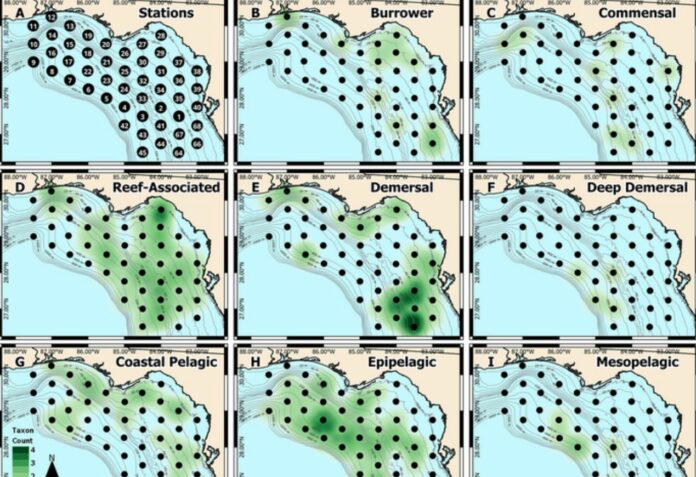
Additionally, the spatial heatmaps of eggs corresponded to known habitat types occupied by adults, indicating that the eggs’ location provides valuable information about their parent species.
The increased throughput allowed by DNA metabarcoding resulted in the identification of taxa not previously detected in this region, suggesting the possibility of episodic spawning events.
Furthermore, the study revealed that metabarcoding can expand the number or geographic range of samples that can be processed, which could lead to more comprehensive data analysis.
However, it’s important to note that one disadvantage of DNA metabarcoding is that it is not quantitative and requires the application of a threshold proportion of sequences to count a taxon as present.
The study emphasizes the significance of establishing long-term monitoring of fish eggs. This is crucial for safeguarding spawning sites and habitats used by fish during their early life stages.
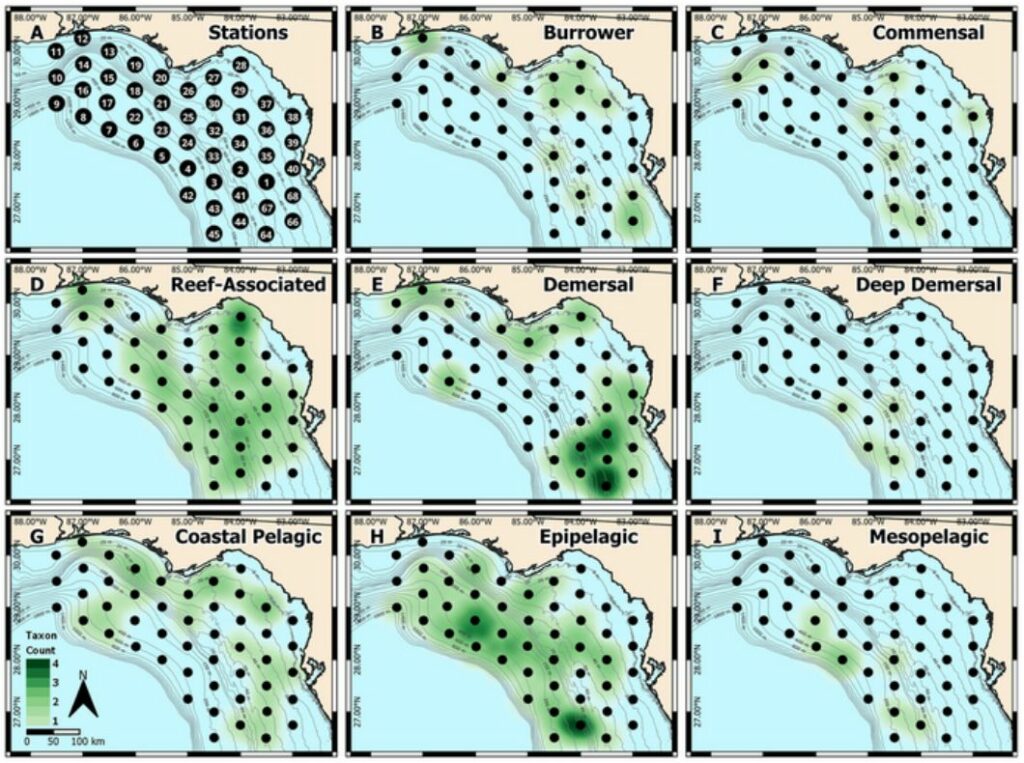
Identifying the composition of fish eggs over an extended period and at high spatial resolution provides valuable insights into changes in the spawning dynamics of various fish species.
Furthermore, such monitoring can help researchers understand how human activities affect fish spawning, making it an essential aspect of conservation and management efforts.
DNA metabarcoding is a technique that amplifies and sequences a conserved gene present in all animals, enabling the identification of the organism by comparing it to a database.
In this particular study, fish eggs were collected using a plankton net, which captures free-floating eggs.
This method is effective since most fish are broadcast spawners and release their eggs into the water column.
The research team processed all the fish eggs together as a composite, significantly reducing the cost of the process.
According to Professor Mya Breitbart from the University of South Florida’s College of Marine Science, the recent study on DNA metabarcoding for fish eggs produced some interesting findings.
They were able to identify eggs from three fish species not previously recorded in the area – Atlantic blue marlin, crested scabbardfish, and burrfish – there were also some unexpected challenges.
In some cases, the number of fish species detected in a sample exceeded the number of eggs present, indicating the presence of contaminating DNA. This likely resulted from “environmental DNA” shed by various living species that can attach to the outside of fish eggs.
To address this issue, the researchers had to apply a threshold for the proportion of sequences belonging to a particular species to determine its presence. However, this approach may have introduced some “biases by excluding rare species.”
According to the researchers, the DNA metabarcoding technique used in the study may not have been the most optimal method for their long-term monitoring goals. However, they believe that there are several potential applications of the technique for fish egg identification.
For instance, the researchers suggest that scientists could collect all the fish eggs from a specific geographic region during a particular season and apply metabarcoding to get a comprehensive overview of fish spawning during that time, which could be compared to data from other seasons.
Source: 10.7717/peerj.15016
Image Credit: Breitbal et al. CC BY 4.0