American geneticists have found that the human hepatitis A virus does not look like other hepatitis viruses but like the “kissing” bugs (subfamily Triatominae) virus. Scientists suggest that bugs were the original owners of the ancestral virus, which then spread to bats, and from them to rodents, monkeys and people.
Human hepatitis can be caused by the reproduction of several viruses, and they all use different breeding strategies. Hepatitis A, C and E viruses contain a sense RNA chain but belong to different families – picornaviruses, flaviviruses and herpes viruses respectively. Hepatitis D virus carries an anti-sense chain of RNA, and the hepatitis B virus belongs to retroviruses. This diversity of the structure suggests that, despite the common target in the human body, these viruses are hardly related to each other.
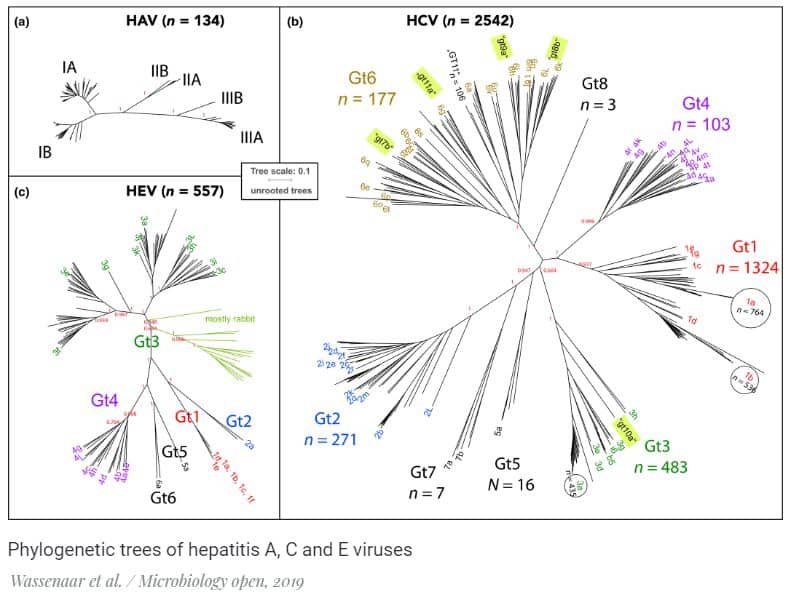
To test this, a team of scientists led by David W. Ussery from the University of Arkansas for Medical Sciences in Arkansas compared the genomes of the hepatitis A, C, and E viruses and their immediate family. The researchers had 134 genomes of different varieties of the hepatitis A virus, 2542 variants of the hepatitis C virus and 557 hepatitis E. The scientists found that hepatitis A viruses are much less diverse than the hepatitis C and E viruses.
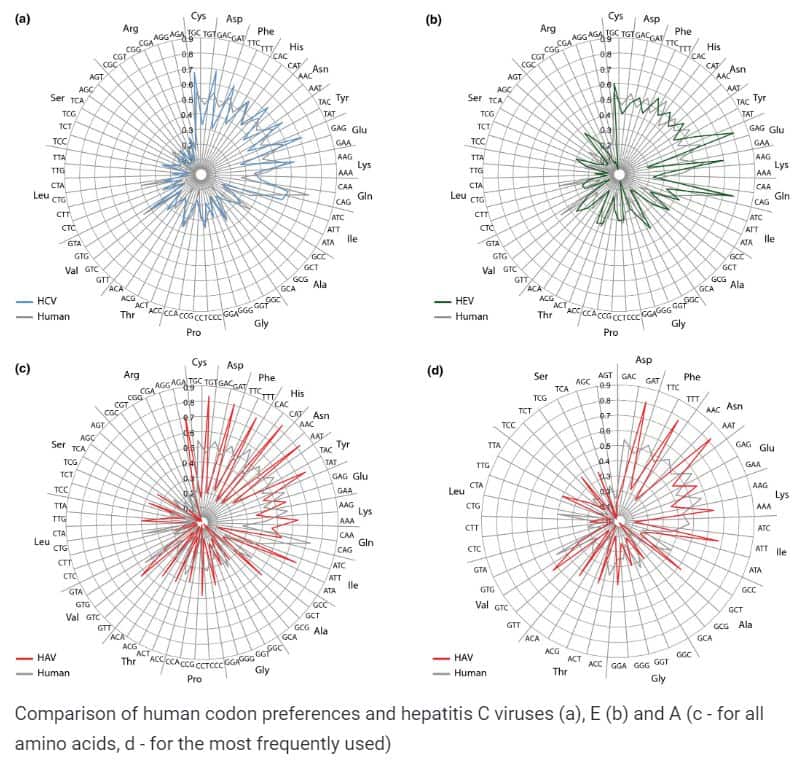
The scientists then analyzed the use of viruses of different codons. Since the genetic code is excess, the same amino acid can be encoded with 1-6 sets of nucleotides, and this trait can also serve as an indicator of kinship: if two organisms or the virus uses a similar set of codons for certain amino acids, you may suspect they have a common origin.
The researchers found that hepatitis A virus differs in the set of codons used not only from hepatitis C and E viruses, but also from its human host. On the one hand, this should be disadvantageous for the virus, because a difference in the genetic code can slow down the synthesis of viral proteins in the host cell. On the other hand, scientists noticed that the differences are rather weak – mainly in the third (last) letter of the codon, to which the cellular protein synthesis machine pays the least attention. Therefore, such a discrepancy in the preferred codons should not seriously affect the success of the viral infection.
The authors of the work suggested that both of these features — low diversity and differences in codon composition — can be explained by the fact that hepatitis A virus came to a person relatively recently, and before that it parasitized on another host with different codon preferences. In hepatitis A viruses that infect other mammals – monkeys, bats and rodents – the set of codons is similar to the human virus, so we can assume that the ancestral virus lived in another group of animals.
In search of a possible host, the researchers first turned to the bacteria that live in the human intestines. They found that Enterococcus faecalis uses a codon-like set of hepatitis A virus. So far, however, not a single bacteriophage (a virus that infects bacteria) with a similar structure has been described, but this may be due to the fact that RNA bacteriophages, in general, are not very well studied.
Then scientists turned their attention to parasitic animals. They noticed that the codon composition of the hepatitis A virus is similar to triatomic bugs – they are also called “kiss bugs” or “kissing bugs” because they often bite people and transmit dangerous diseases, including African trypanosomiasis (sleeping sickness). The structure of the hepatitis A virus was similar to the virus that infects these bugs.
The ancestor of hepatitis A virus and a relative of the modern bug virus, according to the authors, could parasitize on bugs, from them to other mammals – for example, bats and rodents – and from them, it is already possible to reach monkeys and humans. True, so far the virus that would infect insect cells and was similar in genome sequence to the hepatitis A virus has not been described, so scientists will have to find it to confirm their hypothesis.
