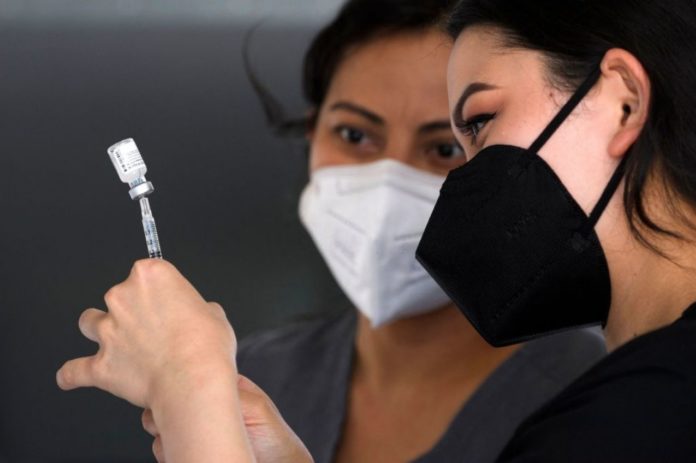
New research shows how SARS-CoV-2 use a variety of methods to escape antibodies produced after an earlier infection with the virus.
The rise of COVID-19 variants of concern – primarily UK B.1.1.7 (also known as the alpha variant) and South African B.1.351 (also known as the beta variant) – has sparked fears that mutations in their spike glycoprotein sequences will influence transmission and result in resistance to multiple neutralising antibodies.
- Scientists in Fear of This New Predator From Red Sea Eating Native Species in Mediterranean
- Does This Mean We Stopped Being Animal and Started Being Human Due to ‘Copy Paste’ Errors?
- The One Lifestyle Choice That Could Reduce Your Heart Disease Risk By More Than 22%
- Aging: This Is What Happens Inside Your Body Right After Exercise
- Immune-Boosting Drink that Mimics Fasting to Reduce Fat – Scientists ‘Were Surprised’ By New Findings
Understanding the specific mechanisms that allow antibodies to escape from SARS-CoV-2 and its strain, which cause COVID-19, is crucial for the development of efficient treatments and vaccines that provide broad protection.
The N501Y mutation, which plays a key role in the interaction between the viral spike glycoprotein and the angiotensin-converting enzyme 2 (ACE2) receptor in human cells, is shared by alpha and beta SARS-CoV-2 variants, according to sequencing findings.
In the research, scientists used wild-type, alpha and beta RBDs to measure the effects of various mutations in that exact region (more specifically, mutations such as N501Y, E484K and K417N) on the binding affinity to ACE2 and a monoclonal neutralizing antibody produced against the original Wuhan-Hu-1 SARS-CoV-2.
Furthermore, this study team used microfluidic antibody affinity profiling to assess the identical wild-type and mutant RBDs. This was done to assess the anti-RBD antibody affinity and concentrations in convalescent serum from people who had been infected with the original SARS-CoV-2 mutation.
The results of the study found that anti-RBD wild-type antibodies are rather effective against wild-type RBD; however, the effectiveness is much lower for both alpha and beta RBD variants, as they follow different approaches to evade the immune response.
More specifically, for RBD-alpha, antibody escape was primarily driven by an increased binding affinity to the ACE2 receptor. In contrast, RBD-beta seems to halt antibody binding by modifying key epitopes on the protein surface.
As a result, both variants show increased effectiveness in binding to ACE2 on the surface of host cells, increasing, in turn, the prospect of successful cell entry. But, strikingly, the antibody concentration binding to RBD-beta was actually half when compared to RBD-alpha and wild-type RBD.
“Our data, therefore, suggest that one factor contributing to the higher transmissibility and antibody evasion of SARS-CoV-2 alpha and beta is a larger fraction of viruses that can form a complex with ACE2”, say study authors. “However, the two variants employ different mechanisms to achieve this goal,” they add.
This study has shown that SARS-CoV-2 alpha RBD can bind to ACE2 with greater affinity, which means the displacement from the receptor by neutralizing antibodies is much more cumbersome; however, RBD-beta is less accessible to antibodies due to epitope changes.
- Scientists in Fear of This New Predator From Red Sea Eating Native Species in Mediterranean
- Does This Mean We Stopped Being Animal and Started Being Human Due to ‘Copy Paste’ Errors?
- The One Lifestyle Choice That Could Reduce Your Heart Disease Risk By More Than 22%
- Aging: This Is What Happens Inside Your Body Right After Exercise
- Immune-Boosting Drink that Mimics Fasting to Reduce Fat – Scientists ‘Were Surprised’ By New Findings
The paper is currently available on the bioRxiv* preprint server while it undergoes a peer-review process.
Photo by PATRICK T. FALLON/AFP via Getty Images