“This is a new way to think about antivirals for RNA viruses”
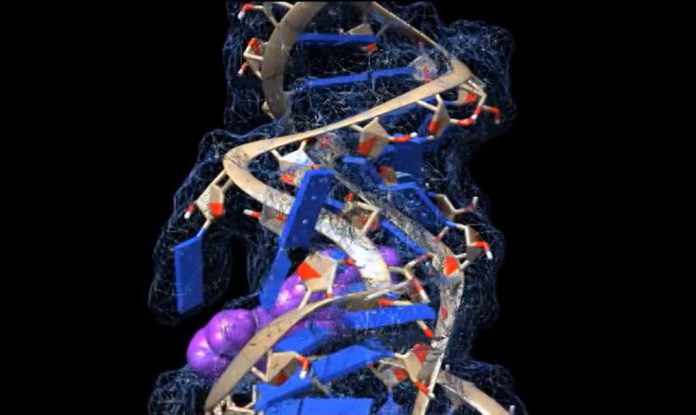
The loops, kinks, and folds in the single strand of RNA that makes up the coronavirus genome resemble a jumble of spaghetti or tangled yarn to the untrained eye.
The unique structures that RNA takes on as it folds into itself, according to experts like Amanda Hargrove, a chemistry professor at Duke University, could have untapped therapeutic promise in the fight against COVID-19.
Hargrove and colleagues report in Science Advances that they have developed chemical compounds that may hook onto these 3D structures and stop virus replication.
“These are the first molecules with antiviral activity that target the virus’s RNA specifically, so it’s a totally new mechanism in that sense,” says Hargrove.
In some regions of the world, the virus is waning, but cases are still rising in others where vaccines are limited. Even in areas where vaccines are widely available, COVID-19 vaccine fear leaves many of the world’s eight billion people vulnerable to infection.
In order to infect your cells, the coronavirus must break in, Coronaviruses enter your cells, send RNA instructions for replication, and hijack your body’s molecular machinery to create more coronavirus replicas. When an infected cell is infected with the virus, it becomes a factory that reads and produces the 30,000 nucleotide “letters” of the virus’s genetic code.
It is these proteins that most antivirals—including the FDA-approved remdesivir, molnupiravir, and Paxlovid for COVID-19—work by binding to. There is a distinct strategy being taken by Hargrove and his colleagues. So far they’ve figured out how to wreak havoc on the viral genome, not only the alphabetical sequence of letters A through U, but the three-dimensional structures that RNA forms.
Hargrove, Blanton Tolbert, Gary Brewer, and Mei-Ling Li from Rutgers were already looking at potential drug options to fight Enterovirus 71, a common cause of hand, foot, and mouth disease in children, when the first scary hints of the pandemic started making news.
Using amilorides, they were able to disrupt the virus’s replication process by binding to hairpin-like folds in the virus’s genetic material.
To investigate if the same compounds could also act against coronaviruses, the researchers first tested 23 amiloride-based molecules against a far less lethal coronavirus that causes many common colds. They discovered three chemicals that, when added to infected monkey cells, lowered the amount of virus in the cells within 24 hours of infection without harming the host cells. They also have stronger effects when given bigger doses. When the compounds were tried on cells infected with SARS-CoV-2, the virus that causes COVID-19, the researchers found comparable outcomes.
Further research revealed that the molecules inhibited the virus’s growth by attaching to a spot in the viral genome’s first 800 letters. The majority of this RNA does not code for proteins, but rather controls their production.
Multiple bulges and hairpin-like formations occur when the region folds in on itself. The researchers were able to study these 3D RNA structures and locate where the chemical compounds were binding using computer modeling and a technique called nuclear magnetic resonance spectroscopy.
The researchers are still trying to find out how these molecules, once attached to the virus’s genome, prevent it from reproducing.
Hargrove believes the field of exploiting RNA as a pharmacological target is still in its early stages. The fact that RNA structures are unstable is one of the reasons. Because they bounce around a lot more than protein cousins, designing compounds that can interact with them in precise ways is difficult.
“The binding pocket that you’re looking for may not even be present most of the time,” Hargrove adds.
Furthermore, 85 percent of the RNA in an infected cell belongs to the human host’s ribosomes, which are biological components made up of RNA and protein.
That doesn’t mean Hargrove isn’t optimistic. For the treatment of spinal muscular atrophy, the FDA has approved a small-molecule medication that acts by attaching directly to non-ribosomal RNA rather than proteins.
On their method, the researchers have a patent application pending. They plan to improve the compounds’ potency before testing them in mice to see if they could be a viable therapeutic option.
The experts believe this isn’t the first time coronaviruses have triggered an outbreak, and it won’t be the last. SARS, which first appeared in China in 2002 and spread to more than two dozen countries, and MERS, which first appeared in Saudi Arabia in 2012, are both caused by the same viral family.
The researchers discovered that the RNA loops and bulges they discovered in bats, rats, and humans, including the ones that triggered the SARS and MERS outbreaks, had stayed virtually unchanged by evolution across related coronaviruses in bats, rats, and humans. That indicates their strategy could be used to combat viruses other than SARS-CoV-2, which produces COVID-19.
Antivirals are clearly valuable weapons to have, so we’ll be better prepared when the next epidemic strikes. Another advantage of having additional medications on hand is the ability to combat resistance. Viruses evolve and change over time. Hargrove believes that being able to mix medications with diverse modes of action will make it less likely for the virus to develop resistance to all of them at the same time and become untreatable.
“This is a new way to think about antivirals for RNA viruses,” Hargrove consludes.
Source: 10.1126/sciadv.abl6096
You were reading: Coronavirus’ twisted RNA strands may offer new treatment options